New: You can test the Web interface to phantom creation for phantomas #betaversion #commentswelcome #contributionswelcome.
Create realistic Diffusion MRI Phantom
Phantomαs is an open-source library written in C/Python designed to create realistic phantoms in structural and diffusion magnetic resonance imaging (MRI). An early version of Phantomαs was used to create the testing and training data of the 2nd HARDI Reconstruction Challenge, organized at ISBI 2013.
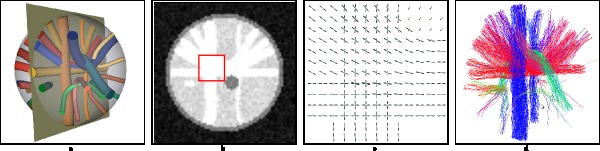
Getting started with Phantomas
Dependencies
Phantomαs is written in Python with parts in C. You will need Python 2.7, and Cython installed on your computer.
Besides, Phantomas depends on:
Optionally, for visualization you may also need:
- VTK (with Python wrapping)
- matplotlib
Build instructions
Phantomas is hosted on GitHub, so first you will need to get the latest version of Phantomas.
To install, simply run the setup.py script. Under linux, simply
run (as root)
# python setup.py install
First steps with Phantomas
Phantomas provides a number of scripts, and examples. The main script is phantomas_dwis. For more info on a script, type:
$ phantomas_dwis -h
Phantomas documentation
Phantomas provides an API to ease integration into your own software or scripts. The API documentation of Phantomas is now available. Phanomas will be presented at the ISMRM 2014 annual meeting; you can download the ISMRM abstract.